BACTERIOPHAGES HELPS TO IMPROVE IMMUNITY DURING MOULTING

Dr Krishna Chandra Sahoo, Global Product Manager, Vetphage Pharmaceutical
In current scenario poultry farmers are struggling to get the availability of maize & soya to feed their flocks due to lock down situations. Feed ingredient prices have soared up, after paying higher prices also they are not getting the ingredients to prepare their poultry feed. So, the farmers left with no choice, to deprive their flocks from feed & putting their flocks on forced moulting irrespective of the ages of birds.
What is forced moulting?
Forced moulting typically involves the removal of food and/ or water from poultry for an extended period of time to reinvigorate egg-laying.
What happens during forced moulting?
During the forced moult, the birds cease producing eggs for at least two weeks, which allows the bird’s reproductive tracts to regress and rejuvenate. Some birds die during forced moulting and it has been recommended that the flock must be well managed so that mortality does not exceed 1.25% over the 1 – 2 weeks of (nearly complete) feed withdrawal, compared to a 0.5% to 1.0% monthly mortality in a well-managed flock under low-stress conditions. Alternative methods of forced moulting which do not use total food withdrawal, e.g. creating a dietary mineral imbalance, generally result in lower mortality rates. After the moult, the hen’s egg production rate usually peaks slightly lower than the previous peak, but egg quality is improved. The purpose of forced moulting is therefore to increase egg production, egg quality, and profitability of flocks in their second or subsequent laying phases.
Consequences of forced moulting?
Forced moulting increases plasma corticosterone which, along with related hormones, decreases the levels of circulating lymphocytes and other leukocytes, thereby reducing the effectiveness of the bird’s immune system. This means that birds being force moulted become susceptible to disease, particularly Salmonella infections, and may produce contaminated eggs thereby compromising public health.
Avian Immune System
Among the avian species, the immune system of the chicken has been studied most extensively. There are many similarities between the general immune mechanisms of mammals and chickens. There are also important differences. Birds respond to antigenic stimulation by generating antibodies as well as cellular immunity. There are three principal classes of antibodies in birds i.e., IgM, IgG (also called IgY) and IgA. Antibody diversity is achieved by gene conversion. T cells are the main effector cells of cellular immunity. The avian T cells differentiate into two distinct pathways i.e., alpha/beta and gamma/delta. Avian T cell diversity is likely generated through combinatorial and junctional mechanisms similar to the mechanisms that operate in mammalian T cell receptors. As in mammals, avian T cells engage in helper and cytotoxic functions that are MHC restricted. The innate effector mechanisms include those mediated by natural killer (NK) cells and antibody dependent cellular cytotoxicity (ADCC). Recently, genes of several avian cytokines have been cloned and expressed. A number of naturally occurring viruses, other pathogens & moulting also cause immunosuppression in chickens. There is much current interest in understanding the mechanisms of immunosuppression and developing strategies to enhance immune responsiveness in commercial poultry. The usage of bacteriophages can help to fight against the pathogens & to improve the immunity.
How phages helps to improve immunity
Phages helps to stimulate immunity through constant interaction of the body with the phages, for example there is continuous stimulation of immune cells to produce interferon gamma. Phages occurrence is common, so that’s why our immune system is all the time stimulated by phage’s antigens.
a. Effects of phages on the host mucosal tissues
Phages interact with host immunity at the mucosal surface. The mucosal surface (e.g., the gut and respiratory tract) represents a critical immunological and physiological barrier that both protects against invading bacterial pathogens while also supporting large communities of commensal microorganisms. The mucosal surface is predominantly composed of mucin glycoproteins that are secreted by the underlying epithelium. By offering both structure and nutrients, mucus layers influence the composition of the microbiota and select for commensal symbionts. It has been shown that mucosal surfaces of the gut commonly support more abundant and stable bacterial populations than the surrounding environments (e.g., the luminal content of the gut). This is, in part, due to the degradation of mucins by gut microbes, but also in part due to host epithelial secretions that selectively shape the commensal microbiota. These host secretions are diverse and can include antimicrobials, such as alpha-defensin and RegIII. Conversely,when mucosal surfaces are invaded by pathogenic bacterial species, the epithelium may respond by increasing the production of antimicrobial agents, hypersecretion of mucin, or alteration of mucin glycosylation patterns in an attempt to subvert microbial attachment and to increase physical removal of the invading bacterial species.
These mucosal layers also harbor large and diverse communities of phages (Fig 1A). Mucus-associated phage communities are significantly enriched compared to the surrounding non-mucosal environment. This increase in phage abundance happens through an adherence mechanism whereby phages weakly bind mucin glycoproteins via immunoglobulin-like (Ig-like) protein domains displayed on their capsids. The Ig-like fold is one of the most common and widely dispersed in nature, present in antibodies and T-cell receptors where it mediates important binding interactions of the host adaptive immune system. These Ig-like domains are found within approximately one quarter of sequenced Caudovirales genomes, and are typically displayed on the virion surface. Most of these structurally displayed Ig-like domains are dispensable for phage growth in the laboratory, which led to the hypothesis that they aid the phage in the adsorption to their bacterial host under environmental conditions. Phages that utilized Ig-like domains, which effectively bind to the mucus layer, would be under positive selection within the mucosa, leading to the proposal of a bacteriophage adherence to mucus (BAM) model as a non-host-derived layer of immunity, mediated by phages .
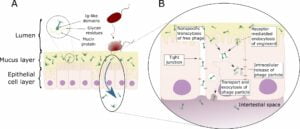 Fig 1. Schematic representation of direct interaction of phages with host epithelial cells.
Fig 1. Schematic representation of direct interaction of phages with host epithelial cells.
(A) Bacteriophage adhering to mucus (BAM). Mucus is produced by the underlying epithelium. Phages of different morphologies can bind variable glycan residues displayed on mucin glycoproteins through variable capsid proteins, such as Ig-like domains. The adherence of phages to this mucus layer creates an antimicrobial layer that reduces bacterial attachment to and colonization of the mucus. This lead, in turn, to a reduction in epithelial cell death. Furthermore, these phages can migrate through theses epithelial cell layers subsequently ending up in the bloodstream.
(B) Phage transcytosis.
Binding interactions between phages and the membrane through transmembrane mucins, specific receptors, or through non-specific recognition, may allow signal transduction in the epithelial cell. Subsequently the phage particle is taken up by the epithelial cell. The internalized phage particles may be degraded leading to intracellular release of phage particles and DNA. Furthermore, it has been hypothesized that phage particles might cross the eukaryotic cell enabling phages to disseminate to the body. Phages may also gain access to the body via a “leaky gut”, where they bypass the epithelial cell barrier at sites of cellular damage or punctured vasculature.
On top of their direct effects on bacterial populations, phages can also have an indirect effect on the colonization of their bacterial host to epithelial cells. In case of Neisseria meningitidis it has been shown that its filamentous phage (MDA) increases its host-cell colonization. The authors showed that the presence of this filamentous phage leads to a higher binding of the bacteria to the host epithelial cells. Furthermore, the phage also seemed to form a linker between the bacteria, further heightening its colonization. These effects were not observed for endothelial cells, indicating a specificity of the phage towards epithelial cells. In this case it is the phage itself that enhances immune system response, promoting bacterial aggregation. It can be further hypothesized whether there is a mutual benefit to phage and bacteria, whereby the phage interacts with the mucosal surface and binds the bacteria. Instead of infecting and lysing the bacteria, the phage would provide the bacteria with additional binding sites, thus, elevating the colonization frequency.
b. Phage Transcytosis
Below the mucosal surfaces, the cellular epithelium forms another physical barrier that separates the heavily colonized mucosa from the normally sterile regions of the body. Due to their ubiquity within the epithelial mucus layer, phages are in constant contact with the epithelial layers. The passage of commensal bacteria colonizing the intestine across the mucosal epithelium to local lymph nodes and internal organs is termed bacterial translocation and is a critical step in the pathology of various disorders. While bacterial translocation is a well-described phenomenon, little is known about the translocation of bacterial viruses.
However, the most important factor seems to be the dose, which correlates strongly with the probability that an orally applied phage can be found in circulation or in tissues. This is in line with the fact that phages may differ in their ability to propagate on gut bacteria and this ability may further limit their systemic dissemination after application per os.
An important consideration regarding the translocation of orally administered phages is whether phages can cross the mucosal barrier in sufficient numbers to subsequently interact with and bypass the cellular epithelium. Recently, it has been demonstrated that phages can enter and cross epithelial cell layers by a non-specific transcytosis mechanism. Phage-epithelial transcytosis seems to preferentially occur in an apical-to-basal direction and was shown to occur across different types of epithelial cell layers (e.g., gut, lung, liver, kidney, and brain cells) and for diverse phage types and morphologies. Microscopy revealed that roughly 10% of epithelial cells endocytosed phage particles, which appeared to be localized within membrane-bound vesicles. Interestingly, those few cells that did endocytose phage particles appeared to contain large numbers of such vesicles. Chemical inhibitor assays suggest that, once endocytosed, phage particles traffic via the Golgi apparatus before being functionally exocytosed at the basal cell layer. The transcytosis of phages across epithelial cell layers provides a mechanistic explanation for the systemic occurrence of phages within the host body in the absence of disease.
Phage Innate Immune Response
a. Phage Phagocytosis
The main function of innate immune cells is the recognition and elimination of foreign material and, when appropriate and necessary, the mounting of an adaptive immune response against it. Neutrophils and granulocytes are critical components of the innate immunity and represent the frontline defense against tissue invasion by bacteria and viruses. Since the 1960s, it has been acknowledged that leukocytes are not only able to bind phages in a time, concentration, and temperature-dependent manner but also internalize and eventually eliminate the phages. Whether specific receptors are involved in phage endocytosis (the term phagocytosis is used for particles >500 nm) is not completely resolved, but recognition of the lysine–glycine–aspartic acid motif in the p26 capsid protein of the T4 phage by β3 integrins on phagocytes might be implicated in this process.
It is well established that phages can be phagocytosed by host cells. As such, the immune system plays a key role in phage clearance from animal and human bodies. Elements of the mononuclear phagocyte system (MPS) in the spleen and liver filter foreign objects, including phages, from the circulation. The spleen and liver have been identified as the major sites of phage accumulation, as phage titers are usually the highest there. The MPS has been credited for the rapid removal of administered wild-type phage from the circulatory system in humans.
Phagocytosis by immune cells within the liver and spleen seem to be the major process of bacteriophage neutralization within the human body. One should note that phagocytosis allows the removal of phage particles, even when no specific response to bacteriophages has been developed. Consequently, phagocytes are probably the major fraction of animal or human cells that interact with bacteriophages in vivo.
Clear evidence concerning the cooperation of phages with the innate immune system was first provided by Tiwari et al., who showed the necessity of a neutrophil-phage cooperation in their solution of P. aeruginosa infections. The authors demonstrated that the presence of neutrophils is necessary to remove phage resistant bacteria, which emerge during the phage therapeutic treatment when only a single phage is used. This was later repeated by Roach et al. and Pincus et al. and converted into an in-silico model by Leung & Weitz.
Studies, in vitro as well as in vivo, regarding the cellular immune response induced by phages have been conducted in recent years and revealed the potential of phages to interact with the mammalian immune system. However, it should be noted that many experiments [concerning the immune response induced by phages have been carried out using phage lysates containing remnants of lysed bacteria (e.g., LPS, cytosolic proteins, or membrane particles) or fragments of the host bacterial cell wall adhered to phage tails. This makes it extremely difficult to determine which components were truly responsible for the modulation of the immune response.
Once activated, macrophages produce an array of microbicide effectors and immunoregulatory cytokines that act in concert to eliminate the invasive agent and influence the course of the developing immune response. Lytic phages induce massive LPS release from Gram-negative bacteria; however, this does not exceed the amounts released under antibiotic treatment. Degradation of phages by polymorphonuclear cells and macrophages has been described for T2, φX174, λ, and P22 phages. These studies demonstrate rapid phagocyte-mediated clearance of phages from organs, in particular the liver, where a rapid decline of highly infective phage titers was observed. Murine Kupffer cells, the resident liver phagocytes, were subsequently shown to be responsible for this process. Furthermore, in the spleen, high titers of phages were detectable irrespective of the route of phage immunization. Here, splenic macrophages were found to mediate clearance of phages, but they displayed an approximately four-times-slower kinetic than those observed with Kupffer cells.
Notably, the degradation process is also a prerequisite for Ag presentation and initiation of adaptive immune responses. It can thus be envisioned that improvement of phage therapy could include selection of phages with natural resistance to phagosomal degradation or rational design of recombinant phages to achieve this. This could avoid or delay induction of phage-specific adaptive immune responses and possibly prolong persistence of phages in immune-competent individuals.
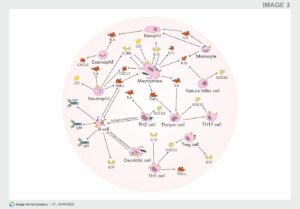
Fig: Interaction of bacteriophages with mammalian immune cells.
Independent of the route of administration, phages can enter the bloodstream and tissues and encounter immune cells in the blood. Phages could encounter these immune cells whilst they are bound to their bacterial host and taken up together by either macrophages or dendritic cells. Alternatively, these phages can directly interact with any of these immune cells by either interacting with cell surface molecules or receptors or taken up using a similar mechanism as observed with phage transcytosis. Once in contact with these immune cells, different pro- (red) or anti-inflammatory (green) cytokines are induced, giving the phage the opportunity to influence the immune response. For example, the induction of IL1RN by the phage blocks the pro-inflammatory signals induced by IL1α and IL1β. Although it is known that phages can induce cytokine response, the precise cells responsible are currently not known. Furthermore, the uptake of phages by antigen presenting cells (APC; e.g., dendritic cells) leads to the activation of B-cells and the production of specific antibodies against the phages.
b. Phage Induced Phagocytosis of Bacteria
Phages can also increase phagocytosis of bacteria by macrophages, since phages administered together with the host bacteria were able to stimulate bacterial phagocytosis. This was attributed to opsonization of bacterial cells by phages, where the phage coats the bacteria and makes it more recognizable for the immune system. This opsonization is in addition to the direct lytic activity of phages, which may contribute to the effective elimination of pathogenic bacteria in vivo. As phages continue the process of infection when adsorbed onto their bacterial host, some authors have suggested that during phagocytosis, phages continue lysing the phagocytosed bacteria, helping the activity of helping the activity of phagocytic cells.
One of the possible responses of phagocytes to foreign objects is the production of reactive oxygen species (ROS). ROS mediate antibacterial activity of phagocytic cells, but excessive ROS production may cause oxidative stress and tissue damage. A preliminary study performed by Przerwa et al. suggested that phage T4 influenced the phagocyte system and inhibited the ROS production in response to pathogenic bacteria (i.e., Escherichia coli). This phenomenon appeared to depend on specific phage-bacterium interactions, but the precise mechanism is currently not known. Furthermore, the host-specific effect could indicate that the ROS reduction is caused by a reduction of bacteria due to infection and lysis by the phage and not due to direct effects by the phage, per se. A possible explanation for a mechanism underlying phage ability to reduce bacteria-induced ROS production in phagocytes was proposed by Miernikiewicz et al., who investigated T4 phage tail adhesin gp12, which specifically binds bacterial LPS and decreased the potency of LPS to induce an inflammatory response in vivo.
c. Cytokine Response against Phages
Several studies have been conducted to determine the potential of phages to induce a cytokine response. Often these studies make use of phage preparations that where not fully purified from bacterial endotoxins or proteins. Several study reports reveal that T7 phage causes inflammatory cytokine production in mice, although no histological changes were observed in the tissues or organs.
In vitro studies have indicated that phages could have anti-inflammatory properties. Recent research studies have also shown clearly that the potential of phages to induce anti-inflammatory properties unrelated to their antibacterial activities.
It should, however, be emphasized that the potential anti-inflammatory or immunosuppressive action of bacteriophages should not be considered as comparable to physiological effects exerted by well-known anti-inflammatory or immunosuppressive drugs. The precise mechanism as to how phages are able to induce (anti-) inflammatory responses is currently not known, although the antimicrobial effect appears to be one of the factors.
Phage Adaptive Immune Response
a. Anti-Phage Antibody Production
Since phages consist of tightly packed DNA or RNA and a protein coat, formed by relatively large number of proteins or repeating protein units, it appears obvious that neutralizing antibodies should be produced in individuals subjected to phage therapy or exposed to naturally occurring phages. Naturally occurring bacteriophages also induce humoral immunity. Phage-neutralizing antibodies against naturally occurring phages (i.e., not therapeutically administered) were detected in the sera of different species (e.g., mice, horse, or human).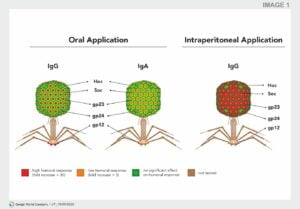
Fig: Antibody induction by phage T4 structural proteins. Individual contribution of T4 head proteins (Hoc, Soc, gp23, gp24, and gp12) to phage immunogenicity. Depending on the administration rote (i.e., oral or intraperitoneal), a difference in antibody response can be observed. When phages are administered orally, strong IgG or low IgA response towards Hoc can be observed, whereas intraperitoneal applications lead to high IgG responses towards Hoc and gp23. Modified Majewska et al.
In-Silico Modeling of the Immune Response Towards Phages
In-silico models predicting phage therapeutic interventions have been developed to better understand the immune response against phages and its impact on the outcome of phage therapeutic interventions. These models are complicated by the fact that phages are protein-based biological agents that interact with the body’s immune system, actively replicate, and even evolve during manufacture or use. As such, phage applications have a vastly different pharmacology compared to conventional drugs. In these mathematical models, the rate at which a bacterial population declines due to phage infection, the rate at which the phage population increases, and the levels at which they are maintained depends primarily on five parameters: the infectivity of the phage, the latency period, the burst size, the rate at which the phages are degraded or removed from the site of infection, and the bacterial growth rate. Besides these five parameters, two other variables need to be taken into account: the density of susceptible bacteria and the density of the phage. In summary, these models describe phage pharmacokinetics as being analogous to the population dynamics of the phage-bacterial interaction, not taking into account potential interaction between bacteria and phages with the innate or adaptive immunity.
These mathematical models can be further extended to include the mammalian host response towards the phage. Based on experimental data, a general scheme can be developed for the tripartite interactions between phage, bacteria, and mammalian immunity. This scheme summarizes the main reciprocal dependencies, specifically the limiting or inducing effects (Fig 1). There are three initial key assumptions on which this scheme is based. First, the innate immunity is activated by the bacteria and acts against the bacteria, but at the same time it also acts against the phage. The second assumption is that phages are not able to boost an innate immune response. The third assumption is that the adaptive immunity specific to phages and the adaptive immunity specific to bacteria have no interfering cross-talk. This led to the development of a model with a set of immunity-representing variables; innate immunity (I), adaptive immunity specific to phages (A), and adaptive immunity specific to bacteria (B). A similar in-silico model described the outcome of a phage therapeutic intervention, taking into account the occurrence of phage resistant bacteria and a phage decay rate, which represents both the innate and adaptive immunity towards the phage. This model indicated that neutrophils are necessary to completely clear a bacterial infection when phage resistance occurs, although it could be argued that phage resistance could partially be prevented when using a phage cocktail.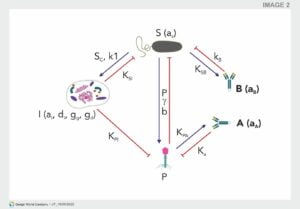
Fig : Schematic representation of the immune response against phages and bacteria. P–Phage, S–bacteria, I–innate immunity, A–adaptive immune response to phage, B–adaptive immune response to bacteria. Green arrows represent a stimulatory effect, red arrows represent an inhibitory effect. Adapted from Hodyra-Stefaniak et al.
When no interaction occurs between the innate immune response (I) and the phage (P), the original Hodyra-Stefaniak model predicts a successful intervention of phages in the removal of a bacterial infection. The inclusion of the variable for the innate immunity (I) demonstrates that the expected outcome of phage therapy could be abrogated by the innate immunity boosted by the bacteria (S) (Hodyra-Stefaniak et al.). Moreover, within the model, the removal of the phage (P) by the innate immune system (I) would lead to a secondary increase in bacterial (S) count, indicating an inefficacy of phage therapy. This is in contrast to the available phage therapy related data. Alternatively, this failure could be counteracted by adjusting the phage dose or changing the timing, as long as the interaction with the innate immunity is considered. Nevertheless, this indicates a shortcoming of the current model described by Hodyra-Stefaniak, indicating further adapting of the model is needed to more closely reflect current knowledge of phage therapeutic outcomes.
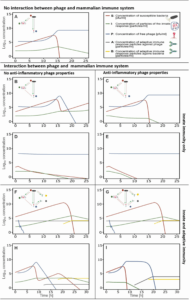
Fig: Effects of innate and adaptive immunity on the success or failure of phage antibacterial treatment, numerical simulations. Innate immune response.
(A) No relation between innate immunity and phage viability. The survival of the phage is independent of the presence of an innate immune response.
(B) Phage susceptibility to the innate immune response. The innate immunity has a negative effect on the phage survival and leads to its removal. Subsequently the bacteria are no longer infected by the phage, and a rise in bacteria is observed.
(C) Phage susceptibility to the innate immune response, considering the anti-inflammatory property of the phage. The anti-inflammatory characteristic of the phage leads to a decline in innate immune particles. This has as effect that the bacterial count diminishes, and the phage survives, similar to A.
(D) Phage susceptibility to innate immune response accommodated and counteracted by an increased phage dose. The higher phage dose leads to the removal of the pathogen and the survival of the phage.
(E) Phage susceptibility to innate immune response accommodated and counteracted by an increased phage dose, considering the anti-inflammatory property of the phage. The effect is the same as in D, but the innate immune response is diminished. Innate and adaptive immune response.
(F) Phage susceptibility to the innate immune response and presence of pre-immunization towards the phage. Presence of pre-existing anti-phage antibodies lead to a rapid drop in phage concentration; hence the phage has no effect on the survival of the bacteria. Once an adaptive immune response towards the bacteria is present, bacterial count decreases.
(G) Phage susceptibility to the innate immune response and no pre-immunization to the phage exists, considering the anti-inflammatory property of the phage. The anti-inflammatory response of the phage has no direct influence on the phage survival in the presence of an adaptive immune response towards the phage. Overall, the response is similar to F.
(H) Phage susceptibility to the innate immune response and no pre-immunization to the phage exists. The absence of a specific adaptive immune response towards the phage leads to a decrease in the bacterial population. The combined effect of innate and adaptive immunity towards the phage leads to a drop-in phage particle concentration.
(I) Phage susceptibility to the innate immune response and no pre-immunization to the phage exists, considering the anti-inflammatory property of the phage. Once the phage reaches a critical concentration (Pc, the concentration of phages needed to induce an anti-inflammatory response), the innate immune response decreases, and the phage concentration grows until all bacteria are removed. Once an adaptive immune response is present against the phage, the phage concentration diminishes until completely removed.
Conclusions
The data reviewed here indicate that phages can interact with the host immune system in a variety of ways that are both direct and indirect. However, the magnitude and nature of the influence that these viruses have on host immunity are only beginning to come into focus. At present, the available data suggest that these interactions tend to be anti-inflammatory. If the observations by Van Belleghem et al. and Zhang et al. concerning the anti-inflammatory properties of phages can be further validated, it is conceivable that phages could influence both our interactions with our commensal flora as well as the outcome of phage therapeutic interventions. Further investigations need to be done, but phages have shown that on administration it helps to improve immunity by various mechanisms.
References
1. Duerkop, B.A.; Hooper, L.V. Resident viruses and their interactions with the immune system. Nat. Immunol. 2013, 14, 654–659. [CrossRef]
2. Proctor, L.; Sechi, S.; Di Giacomo, N.; Fettweis, J.; Jefferson, K.; Strauss, J., III; Rubens, C.; Brooks, J.; Girerd, P.;Huang, B.; et al. The Integrative Human Microbiome Project: Dynamic Analysis of Microbiome-Host Omics Profiles during Periods of Human Health and Disease. Cell Host Microbe 2014, 16, 276–289. [CrossRef]
3. Lloyd-Price, J.; Mahurkar, A.; Rahnavard, G.; Crabtree, J.; Orvis, J.; Hall, A.B.; Brady, A.; Creasy, H.H.; McCracken, C.; Giglio, M.G.; et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature 2017, 550, 61–66. [CrossRef] [PubMed]
4. Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [CrossRef]
5. Lozupone, C.A.; Stombaugh, J.I.; Gordon, J.I.; Jansson, J.K.; Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 2012, 489, 220–230. [CrossRef] [PubMed]
6. Reyes, A.; Semenkovich, N.P.; Whiteson, K.; Rohwer, F.; Gordon, J.I. Going viral: Next-generation sequencing applied to phage populations in the human gut. Nat. Rev. Microbiol. 2012, 10, 607–617. [CrossRef]
7. Oh, J.; Byrd, A.L.; Park, M.; Kong, H.H.; Segre, J.A. Temporal Stability of the Human Skin Microbiome. Cell 2016, 165, 854–866. [CrossRef] [PubMed]
8. Foulongne, V.; Sauvage, V.; Hebert, C.; Dereure, O.; Cheval, J.; Gouilh, M.A.; Pariente, K.; Segondy, M.; Burguière, A.; Manuguerra, J.C.; et al. Human skin Microbiota: High diversity of DNA viruses identified on the human skin by high throughput sequencing. PLoS ONE 2012, 7, e38499. [CrossRef]
9. Pride, D.T.; Salzman, J.; Haynes, M.; Rohwer, F.; Davis-Long, C.; White, R.A.; Loomer, P.; Armitage, G.C.; Relman, D.A. Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J. 2012, 6, 915–926. [CrossRef] Viruses 2019, 11, 10 16 of 22
10. Wang, J.; Gao, Y.; Zhao, F. Phage-bacteria interaction network in human oral microbiome. Environ. Microbiol. 2016, 18, 2143–2158. [CrossRef]
11. Edlund, A.; Santiago-Rodriguez, T.M.; Boehm, T.K.; Pride, D.T. Bacteriophage and their potential roles in the human oral cavity. J. Oral Microbiol. 2015, 7, 1–12. [CrossRef]
12. Dickson, R.P.; Huffnagle, G.B. The Lung Microbiome: New Principles for Respiratory Bacteriology in Health and Disease. PLoS Pathog. 2015, 11, e1004923. [CrossRef]
13. Lim, Y.W.; Schmieder, R.; Haynes, M.; Willner, D.; Furlan, M.; Youle, M.; Abbott, K.; Edwards, R.; Evangelista, J.; Conrad, D.; et al. Metagenomics and metatranscriptomics: Windows on CF-associated viral and microbial communities. J. Cyst. Fibros. 2013, 12, 154–164. [CrossRef] [PubMed]
14. Willner, D.; Haynes, M.R.; Furlan, M.; Hanson, N.; Kirby, B.; Lim, Y.W.; Rainey, P.B.; Schmieder, R.; Youle, M.; Conrad, D.; et al. Case studies of the spatial heterogeneity of DNA viruses in the cystic fibrosis lung. Am. J. Respir. Cell Mol. Biol. 2012, 46, 127–131. [CrossRef]
15. Reyes, A.; Haynes, M.; Hanson, N.; Angly, F.E.; Heath, A.C.; Rohwer, F.; Gordon, J.I. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 2010, 466, 334–338. [CrossRef]
16. Minot, S.; Sinha, R.; Chen, J.; Li, H.; Keilbaugh, S.A.;Wu, G.D.; Lewis, J.D.; Bushman, F.D. The human gut virome: Inter-individual variation and dynamic response to diet. Genome Res. 2011, 21, 1616–1625. [CrossRef] [PubMed]
17. Santiago-Rodriguez, T.M.; Ly, M.; Bonilla, N.; Pride, D.T. The human urine virome in association with urinary tract infections. Front. Microbiol. 2015, 6, 14. [CrossRef]
18. Brüssow, H.; Canchaya, C.; Hardt, W. Phages and the evolution of bacterial pathogens: From genomic rearrangements to lysogenic conversion. Microbiol. Mol. 2004, 68. [CrossRef]
19. Breitbart, M.; Rohwer, F. Method for discovering novel DNA viruses in blood using viral particle selection and shotgun sequencing. Biotechniques 2005, 39, 729–736. [CrossRef]
20. Dinakaran, V.; Rathinavel, A.; Pushpanathan, M.; Sivakumar, R.; Gunasekaran, P.; Rajendhran, J.Elevated levels of circulating DNA in cardiovascular disease patients: Metagenomic profiling of microbiome in the circulation. PLoS ONE 2014, 9, e105221. [CrossRef] [PubMed]
21. Li, S.K.; Leung, R.K.-K.; Guo, H.X.; Wei, J.F.; Wang, J.H.; Kwong, K.T.; Lee, S.S.; Zhang, C.; Tsui, S.K.-W. Detection and identification of plasma bacterial and viral elements in HIV/AIDS patients in comparison to healthy adults. Clin. Microbiol. Infect. 2012, 18, 1126–1133. [CrossRef] [PubMed]
22. Moustafa, A.; Xie, C.; Kirkness, E.; Biggs, W.; Wong, E.; Turpaz, Y.; Bloom, K.; Delwart, E.; Nelson, K.E.; Venter, J.C.; et al. The blood DNA virome in 8000 humans. PLoS Pathog. 2017, 13, e1006292. [CrossRef] [PubMed]
23. Van Belleghem, J.D.; Clement, F.; Merabishvili, M.; Lavigne, R.; Vaneechoutte, M. Pro- and anti-inflammatory responses of peripheral blood mononuclear cells induced by Staphylococcus aureus and Pseudomonas aeruginosa phages. Sci. Rep. 2017, 7, 8004. [CrossRef] [PubMed]
24. Majewska, J.; Beta, W.; Lecion, D.; Hodyra-Stefaniak, K.; Kłopot, A.; Ka´zmierczak, Z.; Miernikiewicz, P.; Piotrowicz, A.; Ciekot, J.; Owczarek, B.; et al. Oral application of T4 phage induces weak antibody production in the gut and in the blood. Viruses 2015, 7, 4783–4799. [CrossRef] [PubMed]
25. Miernikiewicz, P.; Da˛browska, K.; Piotrowicz, A.; Owczarek, B.;Wojas-Turek, J.; Kicielin´ ska, J.; Rossowska, J.; Pajtasz-Piasecka, E.; Hodyra, K.; Macegoniuk, K.; et al. T4 phage and its head surface proteins do not stimulate inflammatory mediator production. PLoS ONE 2013, 8, e71036. [CrossRef] [PubMed]
26. Hodyra-Stefaniak, K.; Miernikiewicz, P.; Drapała, J.; Drab, M.; Jonczyk-Matysiak, E.; Lecion, D.;Kazmierczak, Z.; Beta, W.; Majewska, J.; Harhala, M.; et al. Mammalian Host-Versus-Phage immune response determines phage fate in vivo. Sci. Rep. 2015, 5, 3–8. [CrossRef]
27. Handley, S.A.; Thackray, L.B.; Zhao, G.; Presti, R.; Miller, A.D.; Droit, L.; Abbink, P.; Maxfield, L.F.; Kambal, A.; Duan, E.; et al. Pathogenic simian immunodeficiency virus infection is associated with expansion of the enteric virome. Cell 2012, 151, 253–266. [CrossRef] [PubMed]
28. McDaniel, L.; Breitbart, M.; Mobberley, J.; Long, A.; Haynes, M.; Rohwer, F.; Paul, J.H. Metagenomic analysis of lysogeny in Tampa Bay: Implications for prophage gene expression. PLoS ONE 2008, 3, e3263. [CrossRef]
29. Scarpellini, E.; Ianiro, G.; Attili, F.; Bassanelli, C.; De Santis, A.; Gasbarrini, A. The human gut microbiota and virome: Potential therapeutic implications. Dig. Liver Dis. 2015, 47, 1007–1012. [CrossRef] Viruses 2019, 11, 10 17 of 22
30. Kim, M.-S.; Park, E.-J.; Roh, S.W.; Bae, J.-W. Diversity and abundance of single-stranded DNA viruses in human feces. Appl. Environ. Microbiol. 2011, 77, 8062–8070. [CrossRef]
31. Reyes, A.;Wu, M.; McNulty, N.P.; Rohwer, F.L.; Gordon, J.I. Gnotobiotic mouse model of phage–bacterial host dynamics in the human gut. Proc. Natl. Acad. Sci. USA 2013, 110, 20236–20241. [CrossRef] [PubMed]
32. Breitbart, M.; Haynes, M.; Kelley, S.; Angly, F.; Edwards, R.A.; Felts, B.; Mahaffy, J.M.; Mueller, J.; Nulton, J.; Rayhawk, S.; et al. Viral diversity and dynamics in an infant gut. Res. Microbiol. 2008, 159, 367–373. [CrossRef] [PubMed]
33. Lim, E.S.; Zhou, Y.; Zhao, G.; Bauer, I.K.; Droit, L.; Ndao, I.M.;Warner, B.B.; Tarr, P.I.;Wang, D.; Holtz, L.R. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 2015, 21, 1228–1234. [CrossRef] [PubMed]
34. Rodriguez-Valera, F.; Martin-Cuadrado, A.B.; Rodriguez-Brito, B.; Paši´c, L.; Thingstad, T.F.; Rohwer, F.; Mira, A. Explaining microbial population genomics through phage predation. Nat. Rev. Microbiol. 2009, 7, 828–836. [CrossRef] [PubMed]
35. Enault, F.; Briet, A.; Bouteille, L.; Roux, S.; Sullivan, M.B.; Petit, M.-A. Phages rarely encode antibiotic resistance genes: A cautionary tale for virome analyses. ISME J. 2016, 053025. [CrossRef] [PubMed]
36. Verma, N.K.; Brandt, J.M.; Verma, D.J.; Lindberg, A.A. Molecular characterization of the O-acetyl transferase gene of converting bacteriophage SF6 that adds group antigen 6 to Shigella flexneri. Mol. Microbiol. 1991, 5, 71–75. [CrossRef]
37. Pieraerts, C.; Martin, V.; Jichlinski, P.; Nardelli-Haefliger, D.; Derré, L. Detection of functional antigen-specific T cells from urine of non-muscle invasive bladder cancer patients. Oncoimmunology 2012, 1, 694–698. [CrossRef]
38. Brüssow, H. Bacteriophage-host interaction: From splendid isolation into a messy reality. Curr. Opin. Microbiol. 2013, 16, 500–506. [CrossRef]
39. Davies, M.R.; Broadbent, S.E.; Harris, S.R.; Thomson, N.R.; van derWoude, M.W. Horizontally Acquired Glycosyltransferase Operons Drive Salmonellae Lipopolysaccharide Diversity. PLoS Genet. 2013, 9, e1003568. [CrossRef]
40. Ivanov, I.I.; Frutos R de, L.; Manel, N.; Yoshinaga, K.; Rifkin, D.B.; Sartor, R.B.; Finlay, B.B.; Littman, D.R. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 2008, 4, 337–349. [CrossRef]
41. Mazmanian, S.K.; Cui, H.L.; Tzianabos, A.O.; Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 2005, 122, 107–118. [CrossRef] [PubMed]
42. Przybylski, M.; Borysowski, J.; Jakubowska-Zahorska, R.; Weber-Da˛browska, B.; Górski, A. T4 bacteriophage-mediated inhibition of adsorption and replication of human adenovirus in vitro. Future Microbiol. 2015, 10, 453–460. [CrossRef] [PubMed]
43. Kawai, T.; Akira, S. Toll-like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 2011, 34, 637–650. [CrossRef] [PubMed]
44. Alexopoulos, L.G.; Saez-Rodriguez, J.; Cosgrove, B.D.; Lauffenburger, D.A.; Sorger, P.K. Networks Inferred from Biochemical Data Reveal Profound Differences in Toll-like Receptor and Inflammatory Signaling between Normal and Transformed Hepatocytes. Mol. Cell Proteom. 2010, 9, 1849–1865. [CrossRef] [PubMed]
45. Blasius, A.L.; Beutler, B. Intracellular Toll-like receptors. Immunity 2010, 32, 305–315. [CrossRef] [PubMed]
46. Shi, Z.; Cai, Z.; Sanchez, A.; Zhang, T.; Wen, S.; Wang, J.; Yang, J.; Fu, S.; Zhang, D. A novel toll-like receptor that recognizes vesicular stomatitis virus. J. Biol. Chem. 2011, 286, 4517–4524. [CrossRef] [PubMed]
47. Focà, A.; Liberto, M.C.; Quirino, A.; Marascio, N.; Zicca, E.; Pavia, G. Gut inflammation and immunity: What is the role of the human gut virome? Med. Inflamm. 2015, 2015, 1–7. [CrossRef] [PubMed]
48. Foxman, E.F.; Iwasaki, A. Genome-virome interactions: Examining the role of common viral infections in complex disease. Nat. Rev. Microbiol. 2011, 9, 254–264. [CrossRef] [PubMed]
49. Farrar, M.A.; Schreiber, R.D. The Molecular Cell Biology of Interferon-gamma and its Receptor. Annu. Rev. Immunol. 1993, 11, 571–611. [CrossRef] [PubMed]
50. Linden, S.K.; Sutton, P.; Karlsson, N.G.; Korolik, V.; McGuckin, M.A. Mucins in the mucosal barrier to infection. Mucosal. Immunol. 2008, 1, 183–197. [CrossRef] [PubMed]




